Posted December 11, 2015
Shu-Bing Qian, Ph.D., Cornell University
 Tuberous sclerosis complex (TSC) is a multisystem genetic disorder characterized by multiple benign and debilitating tumors in the brain, kidney, skin, and other vital organs, leading to neurological disorders, such as severe epilepsy, mental retardation, autism, and a variety of other clinical manifestations. There are approximately 40,000 patients with TSC in the United States alone, and an estimated 900,000 worldwide.
Tuberous sclerosis complex (TSC) is a multisystem genetic disorder characterized by multiple benign and debilitating tumors in the brain, kidney, skin, and other vital organs, leading to neurological disorders, such as severe epilepsy, mental retardation, autism, and a variety of other clinical manifestations. There are approximately 40,000 patients with TSC in the United States alone, and an estimated 900,000 worldwide.
TSC is caused by mutations in the TSC1 and TSC2 genes, which encode tumor suppressors that are associated in a complex. Normal TSC1/TSC2 complexes use multiple cues to regulate the pro-growth kinase mTORC1. High mTORC1 activity promotes unregulated cell growth and likely contributes to tumor formation in TSC patients. Clinical trials have demonstrated the therapeutic value of rapamycin for TSC and associated diseases; however, rapamycin is not effectively killing tumor cells. Targeting multiple effectors in the TSC/mTORC1 pathway may provide additional treatment options.
One central mechanism underlying tumor formation in TSC is uncontrolled protein synthesis by hyperactive mTORC1. Funded by a Fiscal Year 2010 (FY10) Exploration - Hypothesis Development Award from the Tuberous Sclerosis Complex Research Program (TSCRP), Dr. Shu-Bing Qian of Cornell University discovered that cells lacking TSC2 have increased protein synthesis, but reduced protein quality. Building on these findings, Dr. Qian received an FY13 Idea Development Award from the TSCRP to investigate whether this imbalanced protein quality and quantity may contribute to TSC pathology.
Dr. Qian recently developed and described a high-resolution ribosome profiling approach that will allow scientists, himself included, to more accurately map start codon positions and corresponding initiation rates to better understand translational control (how cells regulate protein synthesis). Dr. Qian used this approach, called quantitative translation initiation sequencing, to show variations of start codon selection (not just AUG sequences, which are typically the start site for translation) and also highlighted a dynamic range of initiation rates in response to nutrient starvation. Nutrient starvation (such as amino acid starvation) plays a key role in programmed cell death in TSC-deficient cells. Understanding how nutrient starvation effects translational regulation of protein quality and quantity may provide insight into the pathogenesis of TSC disease and its varying phenotypes.
Dr. Qian will use this technique and others to (1) investigate how TSC/mTORC1 negatively regulates the quality of newly synthesized polypeptides, and (2) elucidate translational re-programming in cells with altered mTORC1 signaling, including those lacking TSC2. The success of this project has many implications. Perhaps most importantly, the identification of novel protein products as a result of TSC/mTORC1 dysregulation will provide pivotal information for future TSC therapeutic development.
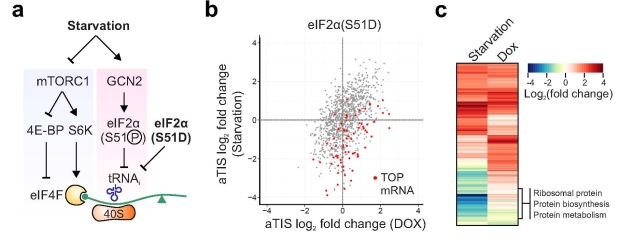
Figure | Distinct role of mTORC1 and eIF2α phosphorylation in translational response to starvation. (a) Schematic of mTORC1 signaling and eIF2α phosphorylation response. (b) Fold changes in LTM-associated aTIS density in eIF2α(S51D) mutant cells between amino acid starvation and DOX-induced eIF2α(S51D) expression. Terminal oligopyrimidine (TOP) mRNAs are shown as red dots. (c) Heat map of fold change of enriched GO terms ("Biological process") for gene groups with translational downregulation (blue) or upregulation (red) in response to amino acid starvation or DOX-induced eIF2α(S51D) expression.
Image credit: Nature Publishing Group via Gao X, et al. Quantitative profiling of initiating ribosomes in vivo. Nat Methods 12(2):147-153.
Publications:
Gao X, Wan J, and Qian SB. 2016. Genome-wide profiling of alternative translation initiation sites. Methods Mol Biol 1358:303-316.
Gao X, Wan J, Liu B, Ma M, Shen B, and Qian SB. 2015. Quantitative profiling of initiating ribosomes in vivo. Nat Methods 12(2):147-153.
Wan J and Qian SB. 2014. TISdb: a database for alternative translation initiation in mammalian cells. Nucleic Acids Res 42(1):D845-850.
Liu B, Han Y and Qian SB. 2013. Co-translational response to proteotoxic stress by elongation pausing of ribosomes. Mol Cell 49(3):453-463.
Conn CS and Qian SB. 2013. Nutrient signaling in protein homeostasis: an increase in quantity at the expense of quality. Sci Signal 6(271):ra24.
Han Y, David A, Liu B, Magadán JG, Bennink JR, Yewdell JW and Qian SB. 2012. Monitoring co-translational protein folding in mammalian cells at codon resolution. Proc Natl Acad Sci 109(31):12467-12472.
Lee S, Liu B, Lee S, Huang SX, Shen B and Qian SB. 2012. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc Natl Acad Sci 109(37):E2424-432.
Links:
Public and Technical Abstracts: Defining Translational Reprogramming in Tuberous Sclerosis Complex
Last updated Thursday, December 5, 2024














