Parkinson's
Understanding the Role of Gene-Environment Interactions in the Degeneration of Human Dopaminergic Neurons in Parkinson's Disease



Posted August 16, 2023
Vikram Khurana, M.D., Ph.D., Department of Neurology, Brigham and Women's Hospital and Harvard Medical School
Beate Ritz, M.D., Ph.D., University of California, Los Angeles
Lee Rubin, Ph.D., Harvard Stem Cell institute
Parkinson's disease (PD) is a degenerative brain disorder, characterized by the loss of dopaminergic neurons, which perform a role in many vital functions in the body, including voluntary movement. A combination of genetic and environmental factors may lead to PD, with genetic factors currently explaining only 10% to 15% of all PD.1 Extensive research on genetic factors has demonstrated that mutations in alpha-synuclein protein and Gaucher protein (GBA) can contribute to PD. Environmental factors that lead to PD are not well understood, but preliminary research has established a correlation between PD and exposure to pesticides.2 Thus, there is a need for research on specific environmental factors and, in particular, the interaction between genes and the environment that contribute to PD.

Vikram Khurana, M.D., Ph.D.
(Photo Provided)

Beate Ritz, M.D., Ph.D.
(Photo Provided)

Lee Rubin, Ph.D.
(Photo Provided)
With a fiscal year 2018 (FY18) Investigator-Initiated Research Award (IIRA), Drs. Khurana, Ritz, and Rubin were funded by the Neurotoxin Exposure Treatment Parkinson's (NETP) program (now managed by Parkinson's Research Program [PRP]) to study the role of gene-environment interactions in the degradation of human dopaminergic neurons in PD. This award was a partnered project, with both Dr. Ritz and Dr. Rubin collaborating with
To complete their study, the team utilized a cohort of participants from a concurrent Pesticide-Wide Association Study (PWAS), separate from the work funded by the IIRA, assessing agricultural pesticide application records from 1974-2017 in Kern, Fresno, and Tulare counties in California. This work used over 1,500 total participants with and without PD. Based on participant residential and occupational proximity to commercial pesticide applications, it was determined that participants living in these agricultural communities sustained exposure to more than 700 pesticides during the 40-year period. The research team excluded pesticides that had fewer than 25 subjects with exposure, resulting in 288 pesticides for the PD-association analysis. Of these, 53 were associated with PD. For the NETP-funded study, the team analyzed data from the PWAS, and results revealed that application of 33 of the PD-associated pesticides were in close proximity to PD patients, including those carrying common mutations in alpha-synuclein protein and GBA.
To understand these effects in the laboratory, the team is assessing the toxicity of the 33 PD-associated pesticides on human dopaminergic neurons. Researchers are using dopamine neurons generated from PD patient induced pluripotent stem cells to test the effects of the various pesticide toxins, providing a personalized disease model in the lab. Significant effort focused on the acquisition, modification, and characterization of multiple patient cell lines. The National Institute of Neurological Disorders and Stroke Human Cell and Data Repository provided human adult stem cell lines, generated from a patient with PD with a mutation in the GBA gene and a control cell line without any mutations, producing a model useful for investigation of the effects of pesticides. With this model, researchers are analyzing how co-exposures to different pesticides can influence the level of toxicity. For example, they have uncovered significant biological effects on dopaminergic neurons of certain pesticides utilized in cotton agriculture. Khurana and his collaborators plan to conduct assays to investigate the mechanisms behind the various toxins. In addition to the in vitro analysis, the team is using the Comparative Toxicogenomics Database, a tool that is focused on understanding how the environment influences health, to identify genetic regions that may increase susceptibility to pesticides.
PD is a debilitating disease, and while genetic factors may not be preventable, environmental factors potentially can be. Dr. Khurana and his team have begun to fill gaps in addressing gene-environment interactions in PD research by compiling epidemiologic data on commonly used pesticides and testing their effects on dopaminergic neurons. Their work will result in quantifiable evidence for the impact of pesticides on PD. Findings from this study could inform regulatory decisions concerning pesticide usage and potentially help generate personalized medicine protocols for patients with PD. The supporting epidemiologic evidence the multi-disciplinary team has collected has built the foundation for future clinical trials. Models created for the PWAS and in vitro analysis will be instrumental resources for further research. Dr. Khurana and his team plan to continue the current study in investigating the effects of pesticide toxins on various patient-derived stem cell lines.
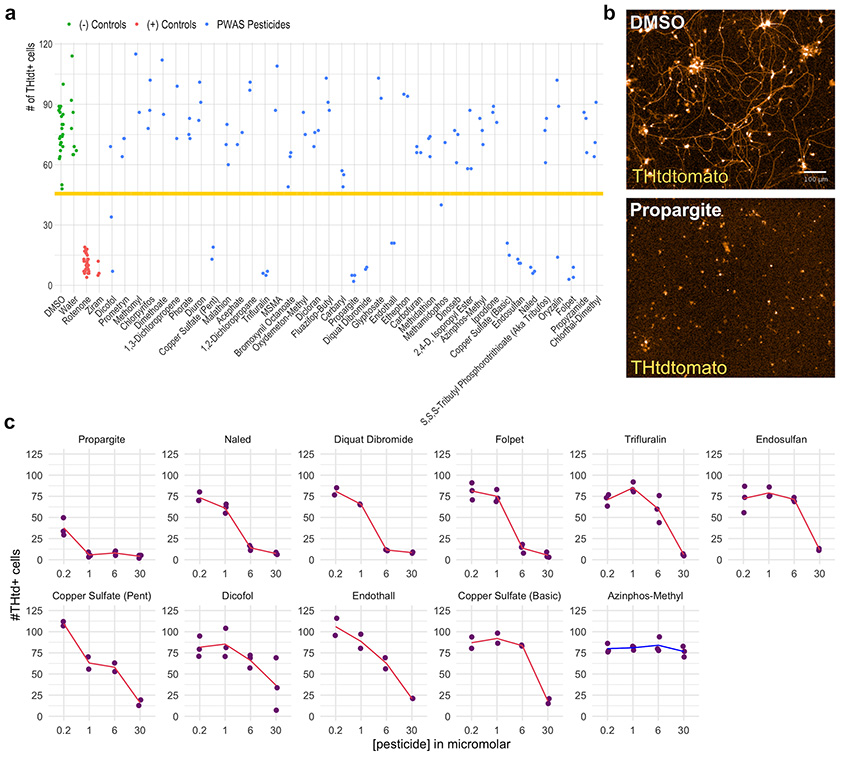 Figure 2. PWAS pesticides are directly toxic to PD patient-derived dopaminergic neurons in a live imaging screen. a) Scatter plot with the number of THtdTomato+ cells measured by live imaging analysis 11 days after the first treatment. DMSO controls (green data points) were present on each assay plate. Water control (green data points) was present on the assay plate containing primarily water-soluble pesticides. Rotenone (red data points) and ziram (red data points) were used as positive controls. Blue data points represent the different pesticides from the PWAS study. Horizontal line denotes three standard deviations below DMSO mean. (b) Upper image is a 10x magnification live image of a DMSO control well. Lower image is from a propargite-treated well. Scale bar = 100µM. More than five independent experiments repeated with DMSO and propargite treatment showed similar neuronal morphology with DMSO and similar extent of cell death and debris with propargite. (c) Four concentration dose curves of PWAS toxicants producing death in SNCA triplication THtdTomato sorted neurons. Cell numbers measured by high content imaging of live cultures 11 days after first treatment. n=1 with two technical replicates per water soluble pesticide and three technical replicates per DMSO and ethanol soluble pesticides per dose per pesticide for the screen dose curves. Red lines connect average cell number value for each pesticide concentration.
Figure 2. PWAS pesticides are directly toxic to PD patient-derived dopaminergic neurons in a live imaging screen. a) Scatter plot with the number of THtdTomato+ cells measured by live imaging analysis 11 days after the first treatment. DMSO controls (green data points) were present on each assay plate. Water control (green data points) was present on the assay plate containing primarily water-soluble pesticides. Rotenone (red data points) and ziram (red data points) were used as positive controls. Blue data points represent the different pesticides from the PWAS study. Horizontal line denotes three standard deviations below DMSO mean. (b) Upper image is a 10x magnification live image of a DMSO control well. Lower image is from a propargite-treated well. Scale bar = 100µM. More than five independent experiments repeated with DMSO and propargite treatment showed similar neuronal morphology with DMSO and similar extent of cell death and debris with propargite. (c) Four concentration dose curves of PWAS toxicants producing death in SNCA triplication THtdTomato sorted neurons. Cell numbers measured by high content imaging of live cultures 11 days after first treatment. n=1 with two technical replicates per water soluble pesticide and three technical replicates per DMSO and ethanol soluble pesticides per dose per pesticide for the screen dose curves. Red lines connect average cell number value for each pesticide concentration.
References:
1Causes. 2022 Parkinson's Foundation, https://www.parkinson.org/understanding-parkinsons/causes .
2Environmental Factors. 2022 Parkinson's Foundation, https://www.parkinson.org/understanding-parkinsons/causes/environmental-factors.
Publication:
Paul KC, Krolewski RC, Moreno EL, et al. 2022. Coupling comprehensive pesticide-wide association study to iPSC dopaminergic screening identifies and classifies
Links:
Project website: khuranalab.bwh.harvard.edu
 An official website of the United States government
An official website of the United States government
 ) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
