Prostate Cancer
LSD1: a novel biomarker and therapeutic target for prostate cancer



Posted February 19, 2019
Shuai Gao, Ph.D.
University of Massachusetts, Boston

Androgen is the male hormone that drives the disease progression of prostate cancer (PCa), and while hormone therapies that target the androgen receptor (AR) are often used to treat PCa, the cancer often stops responding to these hormone therapies and becomes lethal. Therefore, novel therapies are greatly needed. Recent findings from Dr. Changmeng Cai and colleagues while he was at the Beth Israel Medical Center showed that an epigenetic modifier protein called lysine specific demethylase 1 (LSD1) can function broadly as a critical coactivator of AR and could possibly serve as a novel target for PCa therapy. However, much is still unknown about how LSD1 interacts with AR to drive PCa growth.
With support from a FY14 Postdoctoral Training Award, Dr. Shuai Gao at the University of Massachusetts, Boston (UMB) under the mentorship of Dr. Cai (UMB) and Dr. Steven Balk (Beth Israel Medical Center) explored the mechanisms of LSD1 interactions with AR and other co-activators and how these interactions contribute to PCa progression In collaboration with Dr. Housheng Hansen He at the University of Toronto, they demonstrated that LSD1 enzymatic activity is required for the global binding of the FOXA1 protein, and inhibition of LSD1 in vitro globally disrupts FOXA1 binding and AR transcriptional output. Additionally, they identified a site on FOXA1 (lysine 270) that is demethylated by LSD1, which has important implications in mediating AR activity. These results led them to test the effects of introducing an LSD1 inhibitor, GSK2879552, into PCa cell line and patient-derived xenograft (PDX) mouse models that represent castration resistant disease, which showed that the LSD1 inhibitor was able to effectively stop PCa growth.
Overall, these findings add another layer to the interplay between epigenetic modifiers and transcription factors. The positive results from Dr. Gao’s in vivo studies in the PDX model imply strong translational potential of targeting LSD1. The LSD1 inhibitor tested by this research team, GSK2879552, is already in clinical trials for small cell lung cancer, and clinical trials should be launched to investigate the efficacy of LSD1 inhibitors in PCa. In addition, the evidence linking highly methylated FOXA1 and low AR activity suggests methylated FOXA1 as a potential biomarker for patients that may not respond well to hormonal therapy. Recognizing the potential clinical importance of this relationship, this group has already begun developing an antibody to specifically recognize FOXA1 lysine270 methylation that may be incorporated into a diagnostic assay to help provide patients and providers with information to better inform treatment decisions.
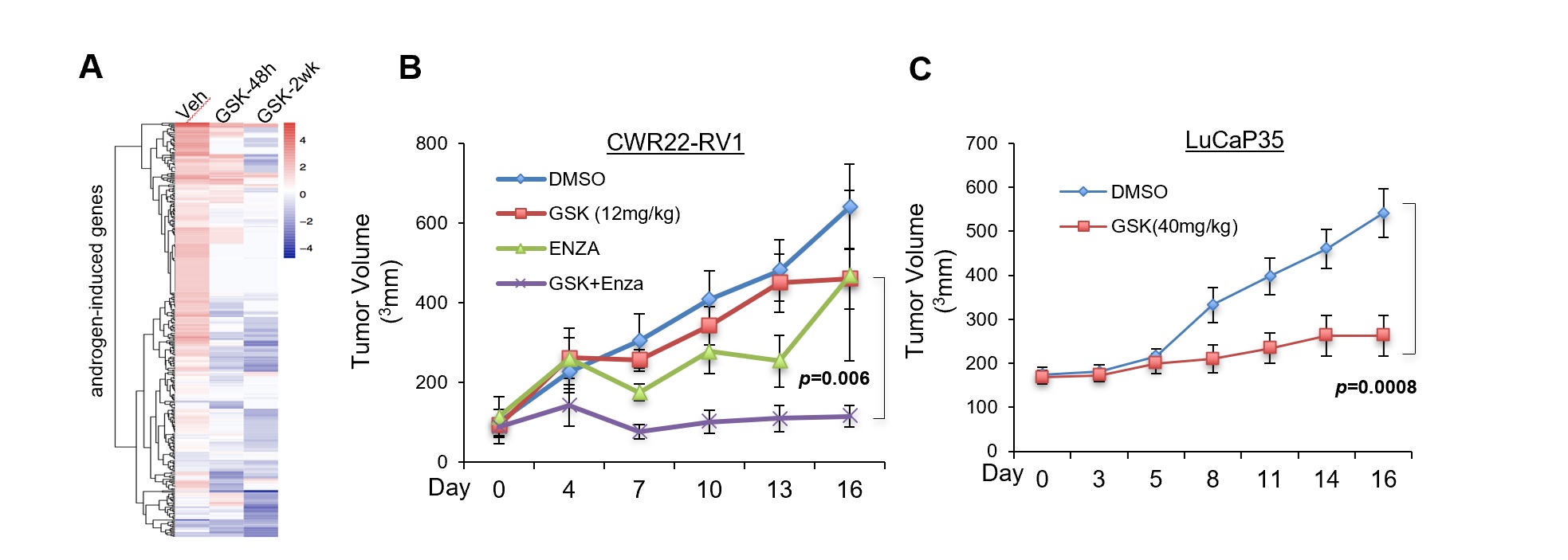
(A) RNA-seq in LNCaP cells stimulated by DHT for 24h and heatmap showing the change in expression of androgen-regulated genes in response to GSK2879552 treatment (1μM). (B) Mice bearing CWR22-RV1 xenograft tumors received daily DMSO, GSK2879552 (12mg/kg), enzalutamide, or GSK2879552 plus enzalutamide via i.p injection, and tumor volume is measured at indicated time. (C) Castrated mice bearing LuCaP35 xenograft tumors received daily DMSO, GSK2879552 (40mg/kg) via i.p injection, and tumor volume is measured at indicated time.
Publications:
Ling, Y., Ahmed, M., Guo, et al. 2017. KDM1A Mediated Epigenetic Reprogramming Drives CENPE Expression and Prostate Cancer Progression. Cancer Res. 77(20):5479-5490.
Han, WT., Gao, S., Barrett, D., et al. 2017.Reactivation of Androgen Receptor-Regulated Lipid Biosynthesis Drives the Progression of Castration-Resistant Prostate Cancer. Oncogene. 2017.385
Cai, C., He, HH., Gao, S., et al. 2014. Lysine-Specific Demethylase 1 Has Dual Functions as a Major Regulator of Androgen Receptor Transcriptional Activity. Cell Reports. 9(5): 1618-27.
Link:
Last updated Thursday, December 5, 2024














