


Posted August 19, 2014
John Kurhanewicz, PhD, University of California at San Francisco
Sabrina Ronen, PhD, University of California at San Francisco
Donna Peehl, PhD, Stanford University
 There is an increasing demand to personalize medicine by tailoring treatment options to each individual patient. It is believed that disease-specific biomarkers can more accurately detect prostate cancer, which would not only improve disease detection and diagnosis, but also measure an individual patient's response to therapy. A team of scientists at the University of California at San Francisco and Stanford University led by Dr. John Kurhanewicz were funded by a fiscal year 2009 PCRP Synergistic Idea Development award to investigate a new molecular imaging technique, called hyperpolarized 13C MRI/MRSI. Hyperpolarized 13C MR techniques exploit the difference in how healthy and cancerous cells metabolize certain chemicals to identify the presence of prostate cancer, its aggressiveness, and its response to therapy.
There is an increasing demand to personalize medicine by tailoring treatment options to each individual patient. It is believed that disease-specific biomarkers can more accurately detect prostate cancer, which would not only improve disease detection and diagnosis, but also measure an individual patient's response to therapy. A team of scientists at the University of California at San Francisco and Stanford University led by Dr. John Kurhanewicz were funded by a fiscal year 2009 PCRP Synergistic Idea Development award to investigate a new molecular imaging technique, called hyperpolarized 13C MRI/MRSI. Hyperpolarized 13C MR techniques exploit the difference in how healthy and cancerous cells metabolize certain chemicals to identify the presence of prostate cancer, its aggressiveness, and its response to therapy.
Critical to their success was the synergistic collaboration between Dr. Kurhanewicz and Dr. Sabrina Ronen, two imaging experts, and an investigator with expertise in prostate cancer and human tissue culture models - Dr. Donna Peehl. Their combined skills enabled them to also develop a MRS-compatible, 3D Tissue Culture Bioreactor for use with primary human prostate tissue cultures (TSCs). This tissue bioreactor provides a true representation of prostate cancer initiation and progression in patients compared to standard animal and cell models, allowing for improved biological identification and interpretation of biomarkers. Using this platform, they identified lactate as a prostate cancer biomarker, and provided the first mechanistic evidence of this marker in living human tissues1,2.
The pre-clinical data generated using this bioreactor helped the investigators secure Investigational New Drug approval from the FDA to continue testing the hyperpolarized 13C MRI in prostate cancer patients. Dr. Kurhanewicz noted from these early clinical trials that "It was gratifying to observe that lactate metabolism in regions of prostate cancer and normal prostate, in the first patient studies, were very similar to what was observed for living normal and malignant prostate tissues inside of the MR compatible 3D tissue culture bioreactor3."
This team of investigators continue to pursue studies to develop companion biomarkers that will track the early development of castrate resistant prostate cancer and its response to therapy. Additional efforts to commercialize the equipment and molecular precursors for performing hyperpolarized 13C MR techniques should bring this technology to patients in the near future.
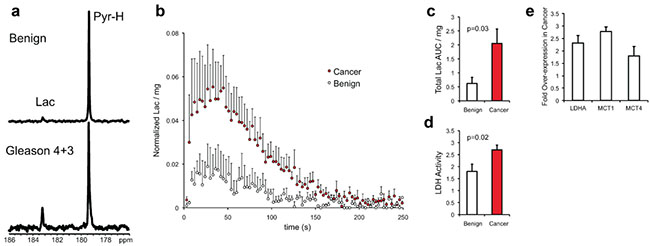
HP [1-13C] Pyruvate to Lactate Flux Is Increasedin Malignant TSCs. a: A single spectrum taken at 90 sec post-injection of HP [1-13C] pyruvate in the bioreactor with benign (N = 4) or malignant TSCs (N = 4). b: Average HP Lactate dynamics for TSCs. c: Total normalized lactate area under the curve (AUC) and (d) LDH activity for be nign and malignant TSCs. e: mRNA expression of lactate dehydrogenase A (LDHA), monocarboxylate transporters1 (MCT1) and 4 (MCT4).
References:
Links:














