Bone Marrow Failure
Novel RNA-Based Aptamer Strategy to Correct Irregular DNA Methylation and Restore Vital Gene Functions in Myelodysplastic Syndromes



Posted September 20, 2023
Annalisa Di Ruscio, Ph.D., Beth Israel Deaconess Medical Center

(Photo Provided)
Myelodysplastic Syndromes (MDS) are a group of diverse bone marrow disorders and the most common acquired bone marrow failure syndrome. They are known to primarily affect the elderly (older than age 65) but could also impact younger patients.3,5 Although not considered a terminal disease at this stage, roughly 30% of those diagnosed with MDS are predicted to progress to acute myeloid leukemia (AML).1,4 DNA undergoes a variety of changes while carrying out its normal functions, but sometimes mistakes occur that can lead to disease; that is what we see in MDS.2
DNA methylation is a normal process that involves the addition of a chemical group, called a methyl group, to DNA, which is the body's way of turning off the activity of a particular gene. Many of the body's functions are regulated by this on/off switch to respond to changing conditions in an appropriate manner. In the case of MDS, errant DNA methylation disables a set of genes called tumor suppressor genes, which silences one of the body's natural protective mechanisms against cancer.2 With funding from a fiscal year 2019 (FY19) Bone Marrow Failure Research Program (BMFRP) Idea Development Award - Early Career Investigator (IDA-ECI), Dr. Annalisa Di Ruscio analyzed factors regulating DNA methylation, with the intent to identify abnormal signaling and ultimately correct it to achieve global and selective DNA demethylation.
Currently, two drugs, Azacytidine and Decitabine, are approved by the U.S. Food and Drug Administration for the treatment of MDS, achieved by demethylating DNA (switching protective gene function back on). However, their clinical applications are limited due to cytotoxic and non-specific demethylation effects, meaning many different genes could be turned on, even some that should stay off. In pursuit of alternatives, Dr. Di Ruscio's team identified a class of RNA that interacts with DNA methyltransferase 1 (DNMT1), a gene that encodes the major enzyme responsible for maintaining methylation patterns following DNA replication. Dr. Di Ruscio discovered that these DNMT1-interacting RNAs (DiRs) block DNMT1 activity, thus preventing DNA methylation (Figure 1). Based on this finding, she and her team developed a RNA-guided tool, in the form of RNA-aptamers (short, single-stranded RNA molecules that can bind to a specific target), to control which genes undergo DNA methylation (Figure 2).
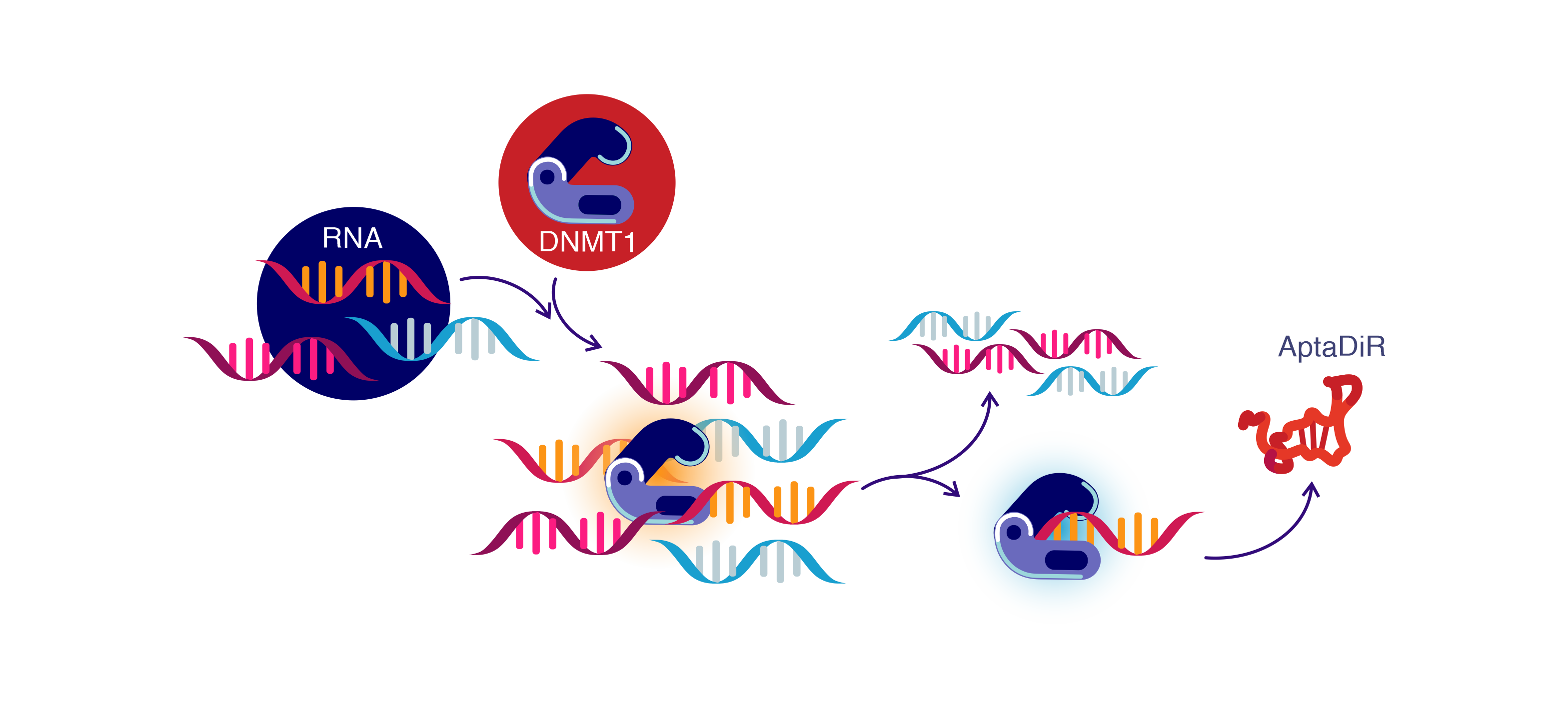 Figure 2: Generation of aptaDiR. Molecules mimicking DiRs - aptaDiRs can be developed to neutralize DNMT1, from a natural existing DiR using an approach called Systematic Evolution of Ligands by EXponential enrichment (SELEX)
Figure 2: Generation of aptaDiR. Molecules mimicking DiRs - aptaDiRs can be developed to neutralize DNMT1, from a natural existing DiR using an approach called Systematic Evolution of Ligands by EXponential enrichment (SELEX)
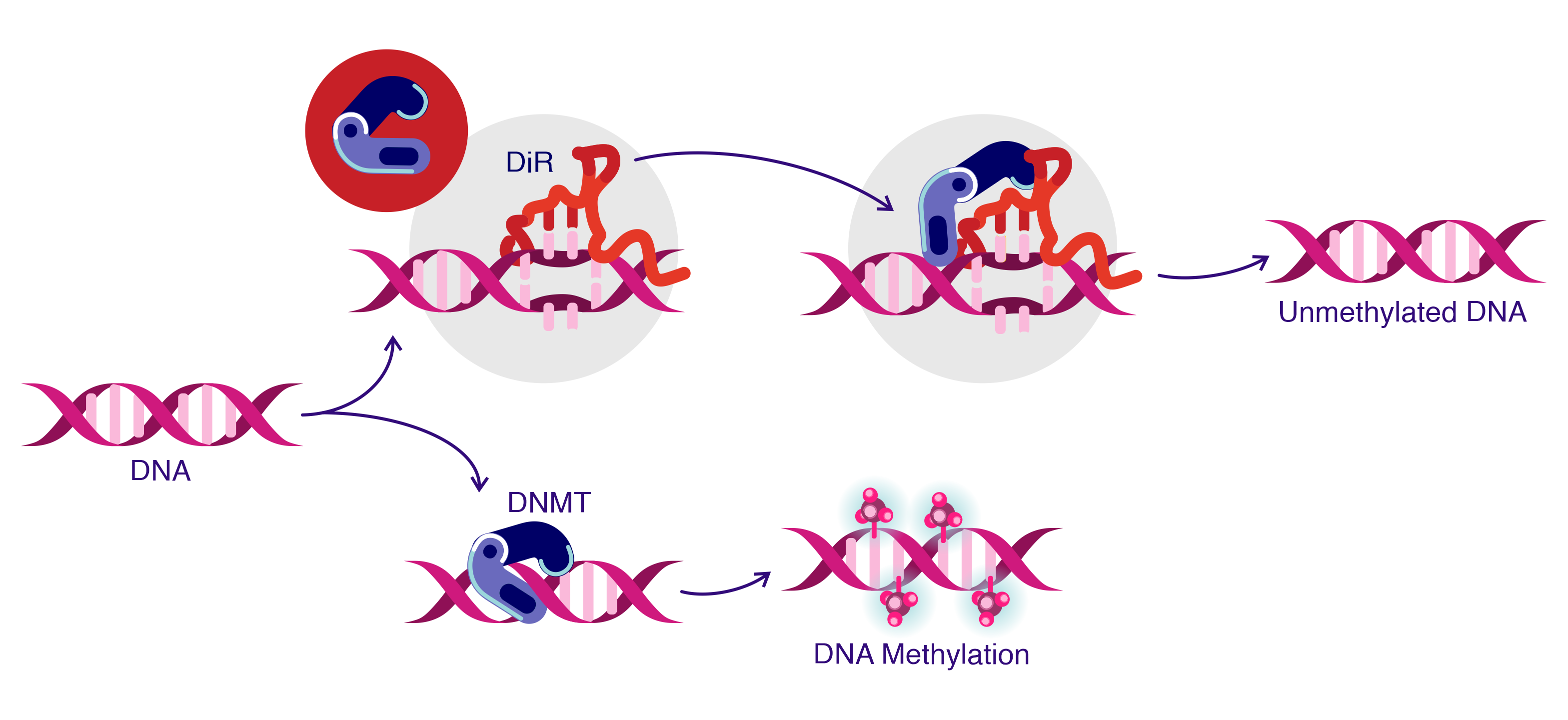 Figure 1: DiR model. DNMT1-interacting RNAs (DiR) inhibit DNMT1 wherein DiRs are present (top panel). In the absence of DiRs, the DNA region can be methylated by DNMT1, (low panel)
Figure 1: DiR model. DNMT1-interacting RNAs (DiR) inhibit DNMT1 wherein DiRs are present (top panel). In the absence of DiRs, the DNA region can be methylated by DNMT1, (low panel)
Using an approach called Systematic Evolution of Ligands by EXponential enrichment (SELEX), Dr. Di Ruscio's team successfully designed several RNA-aptamers. They first verified the aptamers bind securely to DNMT1 with strong affinity. Results from in vitro binding experiments in MDS-like cell lines revealed that two selected aptamers: Ce-49 sh and Ce-10-2 sh, termed aptaDiRs, could bind with high specificity to DNMT1 and displayed increased stability compared to natural existing DiRs. This confirmed their status as promising demethylating agents to act specifically on DNMT1 and achieve reduction in off target effects. Further experiments demonstrated that treatment with Ce-49 sh led to significant demethylation and subsequent reactivation of several tumor suppressor genes (ZCCHC3, PAX5, PLOD2, GAD2, RABA9 and FOXP1), reinstating natural protections against cancer at a rate more potent than that produced by Azacytidine.
These results, taken together with separate studies verifying that aptaDiR Ce-49 sh inhibits progression of human leukemia in in vivo experiments, makes a strong case for the potential utility of these molecules as a treatment strategy for correcting DNA methylation errors in MDS. This data provides a proof of concept for the development of RNA-based therapeutic agents for MDS patients. Dr. Di Ruscio filed a provisional patent application for the DNMT1-specific aptamers and their possible uses. Further, the use of RNA aptamers in regulating DNA expression has far-reaching potential for clinical application in many diseases that involve DNA modifications.
References:
1Arber DA, Orazi A, Hasserjian R, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood . 2016;127(20):2391-2405. doi:10.1182/blood-2016-03-643544
2Figueroa ME, Skrabanek L, Li Y, et al. MDS and secondary AML display unique patterns and abundance of aberrant DNA methylation. Blood . 2009;114(16):3448-3458. doi:10.1182/blood-2009-01-200519
3Jiang Y, Dunbar A, Gondek LP, et al. Aberrant DNA methylation is a dominant mechanism in MDS progression to AML. Blood . 2009;113(6):1315-1325. doi:10.1182/blood-2008-06-163246
4Khan H, Vale C, Bhagat T, Verma A. Role of DNA methylation in the pathogenesis and treatment of myelodysplastic syndromes. Seminars in Hematology 2013;50(1):16-37. doi:10.1053/j.seminhematol.2013.01.001
5Vardiman JW, Thiele J, Arber DA, et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: rationale and important changes. Blood. 2009;114(5):937-951. doi:10.1182/blood-2009-03-209262
Publication:
Esposito CL, Autiero I, Sandomenico A, et al. 2023. Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation. Nature Communications 14(1):99. doi:10.1038/s41467-022-35222-4.
 An official website of the United States government
An official website of the United States government
 ) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
