Neurotoxin Exposure Treatment Parkinson's
The NETP Presents the Fiscal Year 2020 Early-Investigator Research Award Recipients



Posted January 21, 2022
Dr. Deepak K. Gupta, M.D., Larner College of Medicine, University of Vermont
Dr. Kathryn Cross, M.D., Ph.D., University of California, Los Angeles
Dr. Su Jeong Kim, Ph.D., University of Southern California

Dr. Deepak K. Gupta
Dr. Deepak Gupta is a movement disorders neurologist and physician informatician, who has been devoted to Parkinson's disease (PD) since his medical career started. While making several strides in the PD field, driven in major part from personal experiences of supporting his late father Ashok Gupta in living with PD for 13 years, Dr. Gupta is now dedicating his efforts to developing a niche in the application of clinical informatics to advance PD research. With the support from an Early-Investigator Research Award, he plans to develop a multi-institutional clinical informatics research tool, the Ontology-based, Real-time, Machine learning Informatics System for PD (ORMIS-PD), that will be used for predicting individual PD patient's diagnosis probability and prognosis score by using the Movement Disorders Society criteria, open-access PD databases and artificial intelligence methods. The ORMIS-PD tool will assist clinicians and researchers in forecasting two of the most important questions to the patient and their caregivers; how accurate is the patient's diagnosis and how will the patient's disease progress in the future? Dr. Gupta's ORMIS-PD platform from this proposed research project will potentially form the foundation of much needed point-of-care clinical decision support system (CDSS) for PD, and provide new information on potential differences between neurotoxin-associated PD and idiopathic PD in veterans and non-veterans.

Dr. Kathryn Cross
A physician-scientist who has a particular interest to understand the individual differences and heterogeneity in Parkinson's disease (PD), Dr. Kathryn Cross is looking to apply her research experience and training in Movement Disorders neurology in human brain mapping. Specifically, Dr. Cross is looking to study Freezing of Gait (FOG), a heterogeneous, common, and yet poorly understood symptom of advanced PD. With the support from an Early-Investigator Research Award, Dr. Cross plans to unravel phenotypic subtypes of FOG and elucidate dynamics of cortical neural activity throughout the freezing episode. This research study will include PD patients who will wear sensors on the limbs and an electrode cap on their head while neural activity is recorded using electroencephalography (EEG) as they walk in a clinical setting to characterize detailed movement kinematics. This research will potentially increase the understanding of heterogeneity and neurophysiological mechanisms in FOG.

Dr. Su Jeong Kim
Dr. Su Jeong Kim is investigating genetic variants and mitochondrial-derived peptides (MDPs) within Parkinson's disease (PD) that are related to pesticide exposure. Though not everyone exposed to pesticides will develop PD, exposure does have the capability of increasing the risk of PD based on inhibitors and stressors that contribute to mitochondrial dysfunction contained in the pesticide. This research, supported by the Early-Investigator Research Award, seeks to identify the mitochondrial gene-environment interactions (pesticide exposure) and potentially discover novel mitochondrial genetic targets to be used as possible PD treatment targets. Specifically, Dr. Kim is investigating the potential role of SHLP2, a peptide derived from the mitochondria, in determining its therapeutic potential for PD progression. If successful, Dr. Kim's research will create a genomic analysis-based personalized therapeutic approach to treat PD through rapid translation into clinical and population studies.
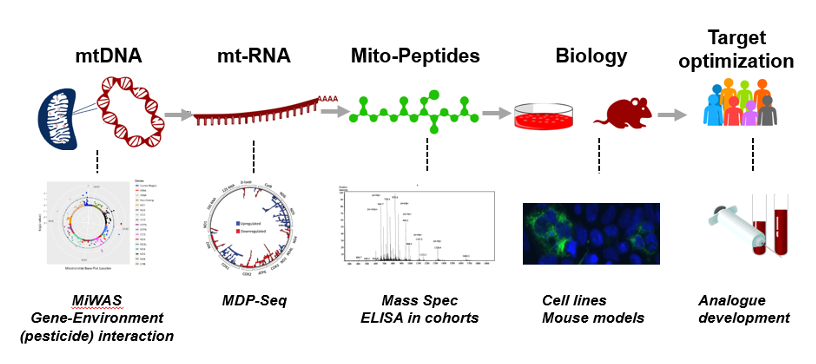
Figure 1: Mitochondrial-derived peptide Discovery Pipeline
Dr. Kim and team applied the systemic approach to discover MDPs and develop the MDP and its analogs as a therapeutic agents. First, they used Mitochondrial Wide Association Studies (MiWAS) that find mutations in mitochondrial DNA sequences that encode for MDPs. Mutations that predict disease provide a novel therapeutic direction for MDPs. They performed MiWAS on Alzheimer's disease, Prostate cancer, NASH (Nonalcoholic steatohepatitis) and Parkinson's disease. Next, the team invented a transcriptome analysis to examine the differential MDP transcript expression in certain diseases and interventions. With the utilization of serum, CSF, and mitochondrial extract they discovered MDPs by mass spectrometry. So far, 41 MDPs were identified. These approaches provide therapeutic direction for MDPs, so they utilized disease relevant in vitro and in vivo models to study the function of MDPs. For the personalized medicine and clinical trials, selecting target patients is the key for success. Optimizing the MDP and its analog based on the genotypes of the patients will be an important step for personalized medicine.
 An official website of the United States government
An official website of the United States government
 ) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
) or https:// means you've safely connected to the .mil website. Share sensitive information only on official, secure websites.
